 Finally, the created network represents the terms as nodes which are linked based on a predefined kappa score level. Since the term–term matrix is of categorical origin, kappa statistic was found to be the most suitable method. Based on this matrix, a term–term similarity matrix is calculated using chance corrected kappa statistics to determine the association strength between the terms. ClueGO creates first a binary gene-term matrix with the selected terms and their associated genes. The relationship between the selected terms is defined based on their shared genes in a similar way as described by Huang et al. An optional redundancy reduction feature (Fusion) assesses GO terms in a parent–child relation sharing similar associated genes and preserves the more representative parent or child term. in certain GO level intervals, with particular evidence codes or with a certain number and percentage of associated genes. Furthermore, the user can adjust the analysis parameters to focus on terms, e.g. To create the annotations network ClueGO provides predefined functional analysis settings ranging from general to very specific ones. To correct the P-values for multiple testing several standard correction methods are proposed (Bonferroni, Bonferroni step-down and Benjamini-Hochberg). Furthermore it provides options to calculate mid- P-values and doubling for two-sided tests to deal with discreetness and conservatism effects as suggested by (Rivals et al., 2007). 2.3 Enrichment testsĬlueGO offers the possibility to calculate enrichment/depletion tests for terms and groups as left-sided (Enrichment), right-sided (Depletion) or two-sided (Enrichment/Depletion) tests based on the hypergeometric distribution. Additionally ClueGO can easily integrate new annotation sources in a plug-in like way ( Supplementary Material). This ensures an up-to-date functional analysis. A one-click update feature automatically downloads the latest ontology and annotation sources and creates new precompiled files that are added to the existing ones. To allow a fast analysis, ClueGO uses precompiled annotation files including GO, KEGG and BioCarta for a wide range of organisms. ClueGO supports several gene identifiers and organisms by default and is easy extendable for additional ones in a plug-in like manner ( Supplementary Material). Gene identifier sets can be directly uploaded in simple text format or interactively derived from gene network graphs visualized in Cytoscape.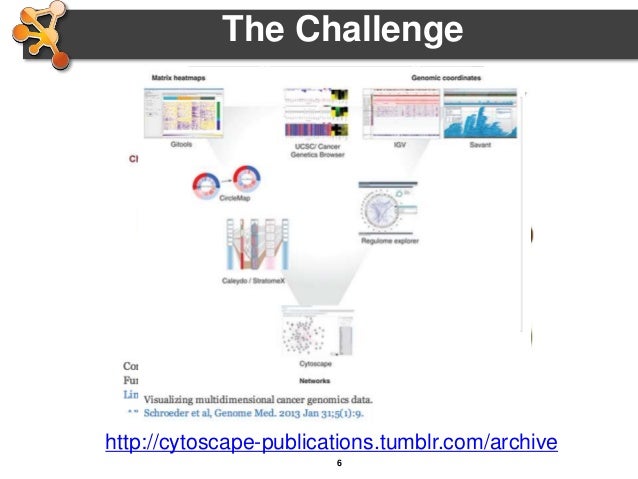 
2 METHODS AND IMPLEMENTATIONĬlueGO has two major features: it can be either used for the visualization of terms corresponding to a list of genes, or the comparison of functional annotations of two clusters. ClueGO takes advantage of Cytoscape's versatile visualization framework and can be used in conjunction with the GOlorize plug-in (Garcia et al., 2007). In addition, ClueGO can compare clusters of genes and visualizes their functional differences. A variety of flexible restriction criteria allow for visualizations in different levels of specificity. ClueGO integrates GO terms as well as KEGG/BioCarta pathways and creates a functionally organized GO/pathway term network. 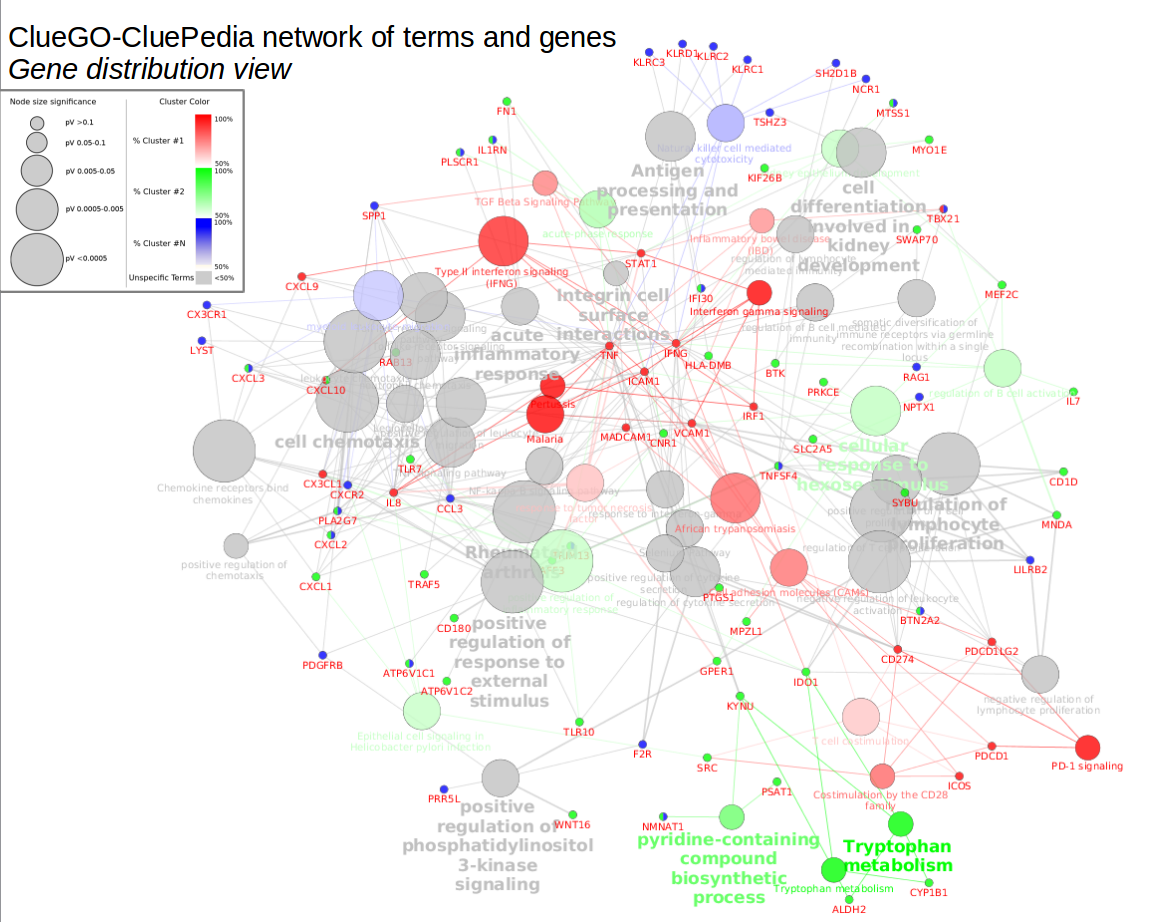
( 2008) which creates an in silico annotation network based on pathways and protein interaction data and maps the gene list of interest afterwards, ClueGO generates a dynamical network structure by already initially considering the gene lists of interest. Compared with the approach of Ramos et al. Other tools like BiNGO (Maere et al., 2005) or PIPE (Ramos et al., 2008) assess overrepresented GO terms and reconstruct the hierarchical ontology tree, whereas ClueGO uses kappa statistics to link the terms in the network. Li et al., 2008) were developed to enhance data interpretation.Īs most of these tools mainly present their results as long lists or complex hierarchical trees, we aimed to develop ClueGO a Cytoscape (Shannon et al., 2003) plug-in to facilitate the biological interpretation and to visualize functionally grouped terms in the form of networks and charts. 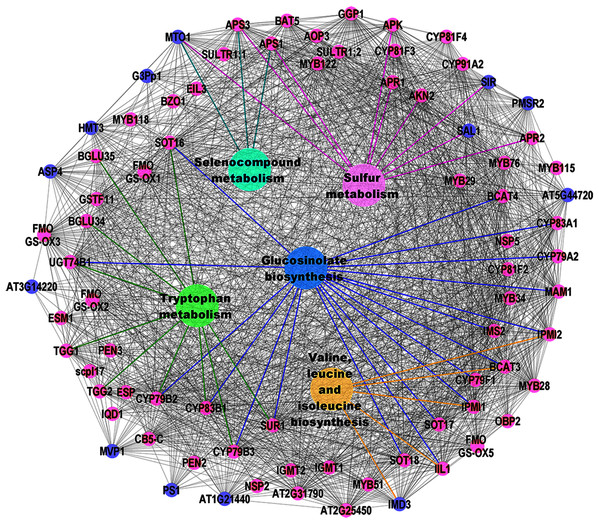
Boyle et al., 2004 Huang et al., 2007 Maere et al., 2005 Ramos et al., 2008 Zeeberg et al., 2003) and algorithms (e.g. Several functional enrichment analysis tools (e.g. For example Gene Ontology (GO) (Ashburner et al., 2000) annotates genes to biological/cellular/molecular terms in a hierarchically structured way, whereas Kyoto encyclopedia of genes and genomes (KEGG) (Kanehisa et al., 2002) and BioCarta assigns genes to functional pathways. Since the number of genes that can be analyzed by high-throughput experiments by far exceeded what can be interpreted by a single person, different attempts have been initiated in order to capture biological information and systematically organize the wealth of data. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
